Apparently, the instrument is very sensitive to external vibrations or even the countertop that it is standing on. 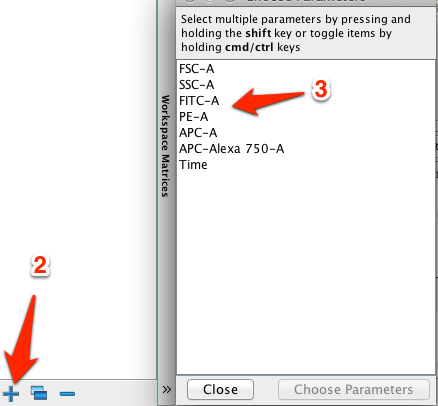
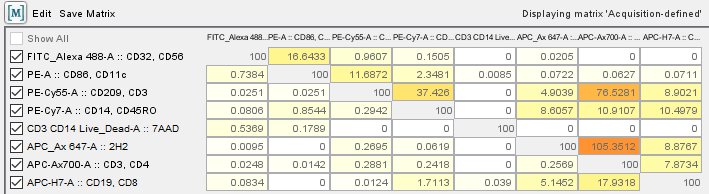
The PMT voltages cannot be changed of course, but the compensation matrix has to be checked.Ĥ. I have been taught that this is because there can be huge differences even if the samples have been acquired with the same template and values. The analysis itself is done in batches but all the compensations are always double-checked beforehand. Thus, tweaking the compensation values is done carefully in order to adjust these discrepancies.ģ. Some samples have a lot of autofluorescence when compared to others. Some compensations look heavily under-/over-compensated. The matrix does not look alike for each sample. Because the compensation depends on the day and the sensitivity of the instrument varies throughout the years of collecting the data, this is a lengthy but crucial step.Ģ. Check the compensation matrix for each sample. This is what I have been taught thus far:ġ. I understand the concept of making correct compensations for the instrument itself, but I am now wondering whether I have been taught wrong about manually adjusting the compensation values in the compensation matrix when analyzing the data in FlowJo. Our lab is currently using BD FACS Verse (8 colors) to immune phenotype primary renal cell carcinoma tumor cells over the span of 5 years. fcs_file_example_import_comp_csv.Hi all, I’m a mere PhD student questioning my very own existence at the moment and in desperate need of some expert advice regarding flow compensations as well as the of tweaking of values in FlowJo.( imported compensation viewed in the Cytobank interface) The more precise value is stored and used behind the scenes. 00 for values that are actually more precise. 01 percent, the values displayed in the interface will be rounded to the nearest. Also, for compensation values with resolution smaller than. Note that the compensation matrix values are displayed as percent instead of decimal. The example CSV attached to this article after upload to Cytobank.  
The example CSV attached to this article before upload: Cytobank also supports the import of values in scientific notation using the letter E (for example, the number 0.00119 written as 1.19E-03). For example, a CSV value of 1 is displayed as 100 in the Cytobank interface.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
